In this discussion thread, feel free to ask questions related to the TB Tracker: WHO package update and lessons from the field session of the 2020 DHIS2 Digital Annual Conference. You can post your questions ahead of, during, or after the session. The panellists will check this thread for questions, and select some for responding to in the session, or follow up after the session has ended. Feel free to respond to other questions or add to them if you have something to follow up with.
Hello all,
Is it possible to give more details about the script (.sh) data transfer tracker-Aggregate?
is it just for TB data or not?
Hi This is for @YuryR does the package has the functions to capture data from GenXpert machines??
Hello @Aanyo! The .sh script is not only for Tb. It can be used for any package.
This is an example of the script (work in progress). Of course, it needs to be modified to meet your parameters.
https://drive.google.com/file/d/1aQ-5rNkRyndS7xmRotshHfDpMlkaqI43/view?usp=sharing
Hi @Zuweina!
At the moment, the Xpert data that is captured in the generic package is
- MTB - detected, not detected, error/invalid/no result
- RIF - susceptible, resistant, indeterminate
- Date of result
- Specimen number
There is no specific function in the package to capture data from GeneXpert machines at the moment. However, we have been discussing this functionality with partners and community. Feel free to get in touch with us so we can collect more information on your use case.
Hi, Community
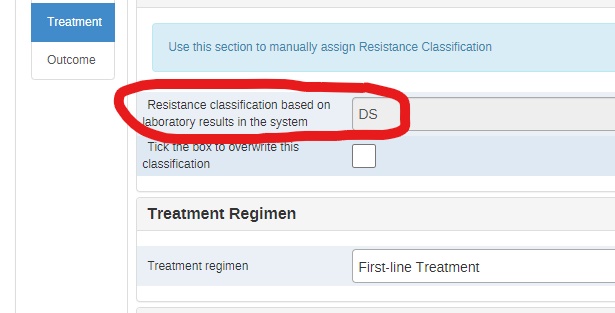
I would like to understand the configuration steps of this field which gives an automatic classification of the resistance according to the laboratory data. this will allow me to adapt the configuration according to the updates of the WHO resistance definitions (Revision of 2021 and 2022).
Note that I have cofigured the WHO TB Tracker 2.36 package.
Thank you for your support.
Hi @elmoujarrade !
Thanks for the question.
To understand the logic of the program I post the link to the design guide here:
If the cases that are registered in DHIS2 are diagnised/confirmed cases, then by default they are registered as DS cases clinically diagnosed.
A whole group of program rule variables, program rule actions and program rules makes sure that as the data is entered into the system, the values get reassigned.
The assignment of resistance classification is based on the laboratory results.
Initial registration == DS
Resistance to any drug == DR
No resistance to Rif and resistance to either Inh, Emb or Pza == Mono res
No resistance to Rif and resistance to a combination of any of the 2 (Inh, Emb, Pza) or all 3 == Poly res
Resistance to Rif only == RR
Resistance to Rif and Inh == MDR
Resistance to Rif, Inh, fluoroquinolones and any 2-line injectables == XDR
No laboratory results + second-line treatment == RR/MDR
The program rules evaluate the resistance to a specific drug entered for any diagnostic test across all diagnostic lab events and assign true values to calculated program rule variables.
Other program rules (with priority configuration) evaluate conditions (PRV values and assign the values to the data element).
Hope that helps!
Hi, @YuryR
Your answer helped me understand the logic behind the design of the program.
Now I will try to understand how the rules of the program have been configured to modify some of them according to the new WHO definitions.
Thanks for your help.